The MetaModel
This page describes the concept of the MetaModel. Although not mandatory, having read the [Accessing and querying the database] documentation may help understand this section.
- What is the MetaModel? (A brief summary with diagrams of what the MetaModel is conceptually)
- What is it useful for? (Provides examples of what the MetaModel can be used for with code examples)
What is the MetaModel?
PickCells organises the data in the form of a property graph (see also ‘PickCells Concepts’).
Below is an example:
Given the following biological example:
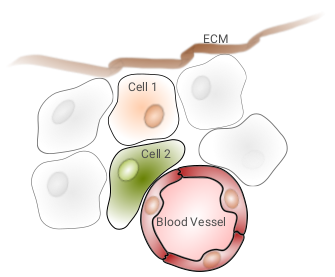
The data graph may look like this:

The MetaModel is another graph which provides information about the data graph. It displays its structure. It can be seen as the ‘schema’ of the data graph. The MetaModel graph for the above data graph may look like this:
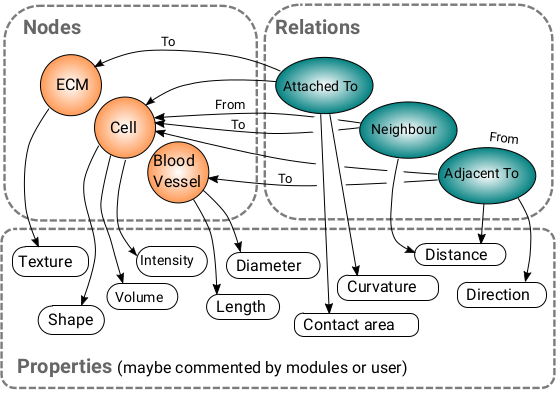
What is it useful for?
The MetaModel provides information about what types of data are present at any given point in the database as well as how they are connected to one another.
In code, MetaModel nodes and links can easily be obtained from a DataAccess instance:
// Get all node types currently present in the database
final Set<MetaClass> metaClasses = access.metaModel().getAll();
Each MetaClass object correspond to a type of node in the data graph. MetaClass objects offers a variety of methods to obtain further information on the object they describe, below are a few examples
// Obtain all the attributes in a given node type:
Stream<MetaReadable> stream = metaClass.getReadables();
Where MetaReadable is a MetaModel object representing an attribute in a type of data node. This provides info about the type of value (numeric, string, array, array length..) or whether the attribute can be found in all data object or only in a subset.
// Get the different link types of the node:
Stream<MetaLink> stream = metaClass.metaLinks()
Where MetaLink are the equivalent of MetaClass but for link data objects.
It is also possible to identify the shortest path between two types of objects:
// Get The shortest path between a root MetaClass and a Target MetaClass using
// some constraints on the type of links and nodes that can be traversed.
final MetaPath path = access.metaModel().shortestPath(root, target, constraints);
MetaClass and MetaLink objects also possess methods to list annotations made by the user such as filtered queries:
// Get the different link types of the node:
Stream<MetaFilter> stream = metaClass.filters()
Where MetaFilters encapsulate the logic for a database query which will target a subset of the objects in the database. This ‘query’ ability is possible because MetaFilter implements DataPointer.
Another important idea is that the MetaModel enables the possibility to interact with objects without knowing their specific type: For example, lets say that we would like to count how many objects exist for some given object type chosen by the user. We could provide a dialog which enables the user to choose one MetaClass among all MetaClasses present in the MetaModel.
Once a MetaClass is chosen, counting the number of object can be done as shown below.
// Count how many object exist in the database for a given object (-1 means no limit)
long count = metaClass.initiateRead(access, -1).count().inOneSet().getAll().run()
This is because, like MetaFilter, MetaClass and MetaLink object also implement DataPointer